About the Center for Bioanalysis
Bioanalysis plays a critical role in life sciences research. Advanced bioanalytical methods and tools have facilitated and accelerated understanding of life processes, evolution of diseases, diagnosis of diseases, optimization of therapies, development of drugs, etc. Establishment of the Center for Bioanalysis (CBA) at the University of Oklahoma (OU) enhances the State's competetiveness and ability to retain high-quality researchers.
Goals for CBA
The major goals of CBA are to promote bioresearch, and to foster high-quality bio-scientists. CBA provides a platform for its members to collaborate on transformative research projects, generate preliminary results, and develop multi-PI (principal-investigator) grant applications. CBA also develops education/training programs to foster OU students to suit the increasing demand for high-quality bio-researchers.
Location and Facilities
CBA is located in the west wing of the first floor of Stephenson Life Science Research Center (SLSRC) at 101 Stephenson Parkway, Norman, Oklahoma 73019. Currently, CBA has Mass Spectrometric Facilities (with five mass spectrometers, UPLC-MS, GC-MS, etc.) and a Microfabrication Facility (photomask aligner, wetbench, spin-rinse drier, photoresist spin coater, profilometer, dicing saw, lapping machine, bonding oven, etc.) for micro- and nano-fabrication of glass chips. It also includes many capillary-based DNA and protein analyzers, cell culture facilities, two-dimensional gel electrophoresis systems, laser-induced fluorescence detectors, electrochemical impedance spectrometer, inverted fluorescence microscope with scientific grade CCD camera, etc.
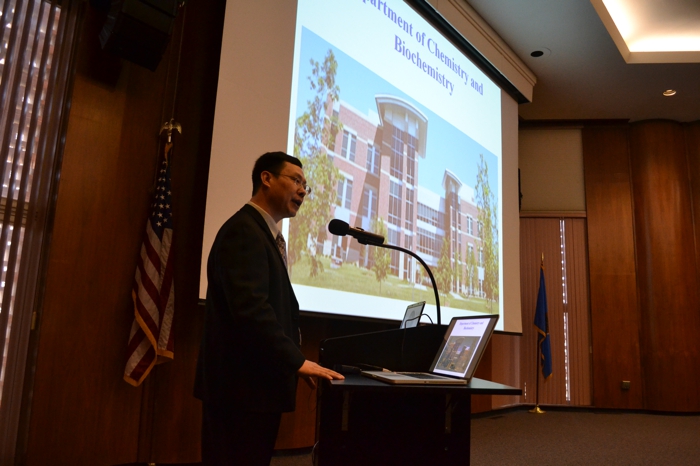
Center for Bioanalysis Sympossium
Research Experiences for Teachers
Strives to involve rural teachers and college instructors in bioanalytical engineering research, improve STEM teaching , and participate in a quality professional network.
Newest CBA Member: Si Wu

Proteomics and Bio-Analytical Mass Spectrometry Group
Bottom-up proteomics has been significantly advanced in last decade, enabling the detection of thousands peptides in a single analysis. However, various protein processing events such as post-translational modifications (PTMs) are frequently untraceable using these approaches. The alternative approach is “top-down” proteomics, where the intact protein samples are directly analyzed by MS, can more accurately reflect protein biological significances and properties of actual biological systems. Our research aims to develop next generation top-down proteomics approaches that combine biochemistry, separation science, mass spectrometer (MS) instrumentation, and bioinformatics to understand protein dynamics in different biological systems (bacteria, fungi, and mammalian cells). Specifically, we are focused on the following directions:
Fungal secretome analysis
Functional characterization of fungi secretome is essential for the understanding of the biochemical regulation and mechanisms involved in the secondary metabolite biosynthesis. Almost all of fungal extracellular enzymes are post-translational modified (e.g., glycosylated). The major objective of our research is to develop a multiplexed PTM analysis platform for characterizing and quantifying the functional proteins constituting the fungal secretome.