
Griswold, Charles M., Jamie Roebuck, Ryan O. Andersen, Lynn F. Stam and Eric P. Spana. 2002. 2002. A toolkit for transformation and mutagenesis in Drosophila using piggyBac. Dros. Inf. Serv. 85: 129-132.
|
|
|
|||
|
|
||||
A toolkit for transformation and mutagenesis in Drosophila using piggyBac.
Griswold, Charles M., Jamie Roebuck, Ryan O. Andersen, Lynn F. Stam and Eric P. Spana. Syngenta Biotechnology, Inc. 3054 Cornwallis Road, Research Triangle Park, NC 27709
We
have generated a high efficiency germline transformation system using the
piggyBac transposable element and a series of vectors for gene misexpression
under the control of the GAL4/UAS system.
We have adapted Gateway recombination technology to the rapid and simple
production of inverted repeat constructs which produce an RNAi effect when
expressed. Finally, we have created
a stable source of piggyBac transposase in the genome that is capable of mobilizing
piggyBac elements in somatic and germline cells.
High efficiency transformation
using ubiquitin-driven transposase
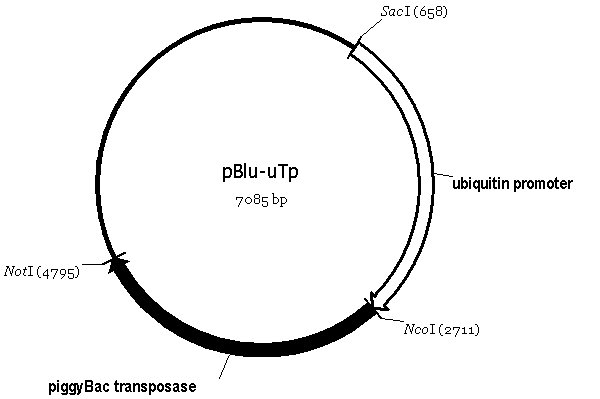
Using the known sequence of the piggyBac transposable element as a reference, we amplified the intact, full-length transposon from genomic DNA of the lepidopteran Trichoplusia ni. Our clone contains one open reading frame with 99.8% identity to that previously described (Handler and Harrell II, 1999). We subcloned this open reading frame via PCR downstream of the poly-ubiquitin promoter (Lee et al., 1988) in pBluescript KS to create pBlu-uTp (Figure 1A). This helper plasmid was then used as the transposase source for the germline transformation of Drosophila. Using 20 different piggyBac vector constructs at a concentration of 600 mg/ml, and the helper plasmid at 400 mg/ml, we achieved transformation rates ranging from 4% to 36% of fertile G0s. This compares favorably to efficiencies obtained by Li et al. (2001) and Handler and Harrell II (1999). Table 1 details the rates of transformation using this helper plasmid.
|
Useful vectors for germline
transformation
The
utility of the piggyBac transformation system is currently limited by the
available plasmid constructs. To
remedy this, we generated four vectors that may prove useful for misexpression.
All of the vectors contain a mini-white transgene at the 5’ end
of the modified piggyBac as the selectable marker.
Mini-white was amplified from flies containing a pUAS P element transgene,
and the integrity of the amplified segment was verified by sequencing. The 5’ and 3’ piggyBac ends
were amplified from
 |
 |
|
A
|
B
|
 |
 |
|
C
|
D
|
 |
 |
|
E
|
F
|
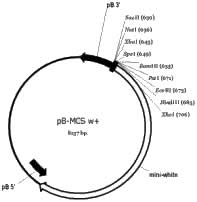
the original Trichoplusia fragment, and cloned into opposite ends of the pBluescript KS multiple cloning site. The mini-white gene was cloned next to the 5’ piggyBac end to generate the plasmid pB-MCS w+ (Figure 1B). This plasmid contains nine unique restriction sites within the transposable element.
Using pB-MCS w+ as a starting point, we generated a vector capable of misexpressing genes using the GAL4/UAS system, similar to Brand and Perrimon (1993). A fragment containing UAS binding sites and the Hsp70 transcriptional start site was amplified from flies harboring a transgene derived from the pUAS P element. The fragment was verified by sequencing and was placed at one end of the pB-MCS w+ cloning site. A fragment containing the SV40 polyadenylation signal was inserted downstream of the pB-MCS w+ multiple cloning site to generate the plasmid pB-UAS w+ (Figure 1C). This arrangement allows genes to be cloned into any of six unique restriction sites under the control of the UAS enhancer. We have demonstrated that genes cloned into pB-UAS w+ and transformed into flies are controlled as expected by the expression of GAL4.
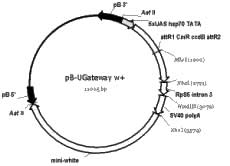
The Gateway cloning technology (Invitrogen), based on lambda phage site-specific recombination, is a rapid and simple method for transferring fragments of DNA between vectors in a directional fashion. We have adapted this technology to two piggyBac vectors to enable cloning DNA fragments into a transformation vector without the use of restriction enzymes. The first vector is based on the pB-UAS w+ plasmid and has a Gateway cassette in a “forward” orientation. Genes transferred from Invitrogen pENTR- or pDONR-derived plasmids will be under the control of the UAS enhancer. A map of this plasmid, called pB-UGateway w+, is shown in Figure 1D.
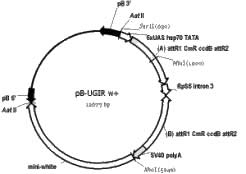
The second vector taking advantage of Gateway cloning technology is called pB-UGIR w+ (Figure 1E). This vector contains a Gateway cloning cassette in the forward orientation just downstream of the UAS binding sites, a 303 bp intron from the RpS5 locus, followed by another Gateway cassette in the reverse orientation. The recombination reaction forces the fragment of interest into both Gateway cassettes and maintains the forward and reverse orientations. Plasmids containing an intact Gateway cassette must be propagated in E. coli containing the gyrA462 allele, such as DB3.1 (Invitrogen). After the recombination reaction (carried out as directed by Invitrogen), the plasmids are transformed into the E. coli strain Stbl2. Only plasmids that have undergone recombination to remove both Gateway cassettes are propagated. We found that all constructs tested were stable in Stbl2 cells (n = 16). Once transformed into Drosophila, transcription initiated by the UAS produces a hairpin RNA (after the intron is spliced out) that serves as a source of dsRNA used in the degradation of endogenous mRNA via RNAi. This vector allows production of an inducible dsRNA by a simple recombinase reaction followed by germline transformation.
Stably integrated Transposase
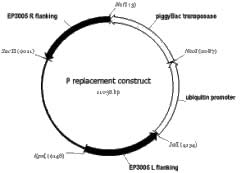
The availability of a stable source of P element transposase in a fly stock has made a wide variety of techniques available for Drosophila research. While such a stock might exist for piggyBac, it is not available from public stock centers. The high transformation frequency of the ubiquitin-transposase plasmid led us to initiate integration of that construct into the Drosophila genome. We used the P element replacement method (Keeler et al., 1996) so that the final product would be stable in any genetic background. We replaced the P element insertion EP3005, located at 100D2. We chose this P element because it was approximately 5 kb from the nearest predicted gene, and showed a high degree of mosaicism when crossed to Delta2-3 stocks. We injected 5292 embryos from a cross between w; EP3005 / TM3, Sb males and y1, w1; Delta2-3, Ki virgins with a replacement construct that carried 2 kb fragments of genomic DNA flanking the EP3005 insert on both sides of the ubiquitin-transposase (Figure 1F). From these injections we collected 336 males of the genotype y1, w1; EP3005 / Delta2-3, Ki. We crossed these males to w; TM3 Sb / TM6B and isolated 375 lines where the w+ of the EP3005 insert had been lost. We established stable stocks from three individuals of each of these lines and screened these 1125 lines by PCR for insertion of the transposase. We isolated one positive, line 36C. Further analysis showed that the ubiquitin-transposase construct was functional and caused the mobilization of w+ piggyBac inserts in the eye. To enhance the genetic usefulness of this insert, we recombined dominant markers onto the 36C insert chromosome. Two stocks are now available: w1118; Tb1, 36C / TM3, Sb and w1118; Sb1, Gl1, Tb1, 36C/ TM3, Ser.
We
tested the ability of the 36C transposase transgene to mediate the transposition
of piggyBac inserts in the germline by examining the mobilization frequency
of two independent inserts. The
first contained only the w+ marker inside piggyBac ends and was
located on a second chromosome marked with Tft1. We could not detect any mobilization of
this piggyBac insertion onto a different chromosome in 55 crosses. The second insertion, a pB-UAS w+ transgene
on the first chromosome, was more amenable. We identified two autosomal jumps from 32 crosses. The percentages shown with these two attempts
indicate that the 36C insert does not catalyze movement of piggyBac elements
frequently enough for mutagenesis screens. It is sufficient, however, to generate new inserts of piggyBac
transgenes already in the genome.
| GenBank accession numbers: | |
| pBlu-uTp | AY196821 |
| pB-MCS w+ | AY196822 |
| pB-UAS w+ | AY196823 |
| pB-UGateway w+ | AY196824 |
| pB-UGIR w+ | AY196825 |
| P replacement construct | AY196826 |